JBrowse2 Genome Browser
Visualize 22 genome assemblies and associated synteny, TE, variant, and multi-omics tracks in JBrowse2.
Data Coverage
- 22 genomes of B. napus with reference sequences (FASTA) and gene annotation.
- Transposable elements (TEs): we integrated 1,058,755 TEs annotated by the EDTA pipeline across 22 genomes.
- ZS11 variation: SNP/InDel variation information derived from 505 resequencing accessions.
- ZS11v1 multi-omics: the platform re-analyzed and aggregated RNA-seq, ATAC-seq, and ChIP-seq tracks for integrative exploration.
- Comparative synteny (collinearity): minimap2 PAF-based tracks across key assemblies (ZS11 and Westar as anchors).Synteny is filtered: alignments shorter than 50 kb are removed to focus on robust collinear blocks.
Quick Start
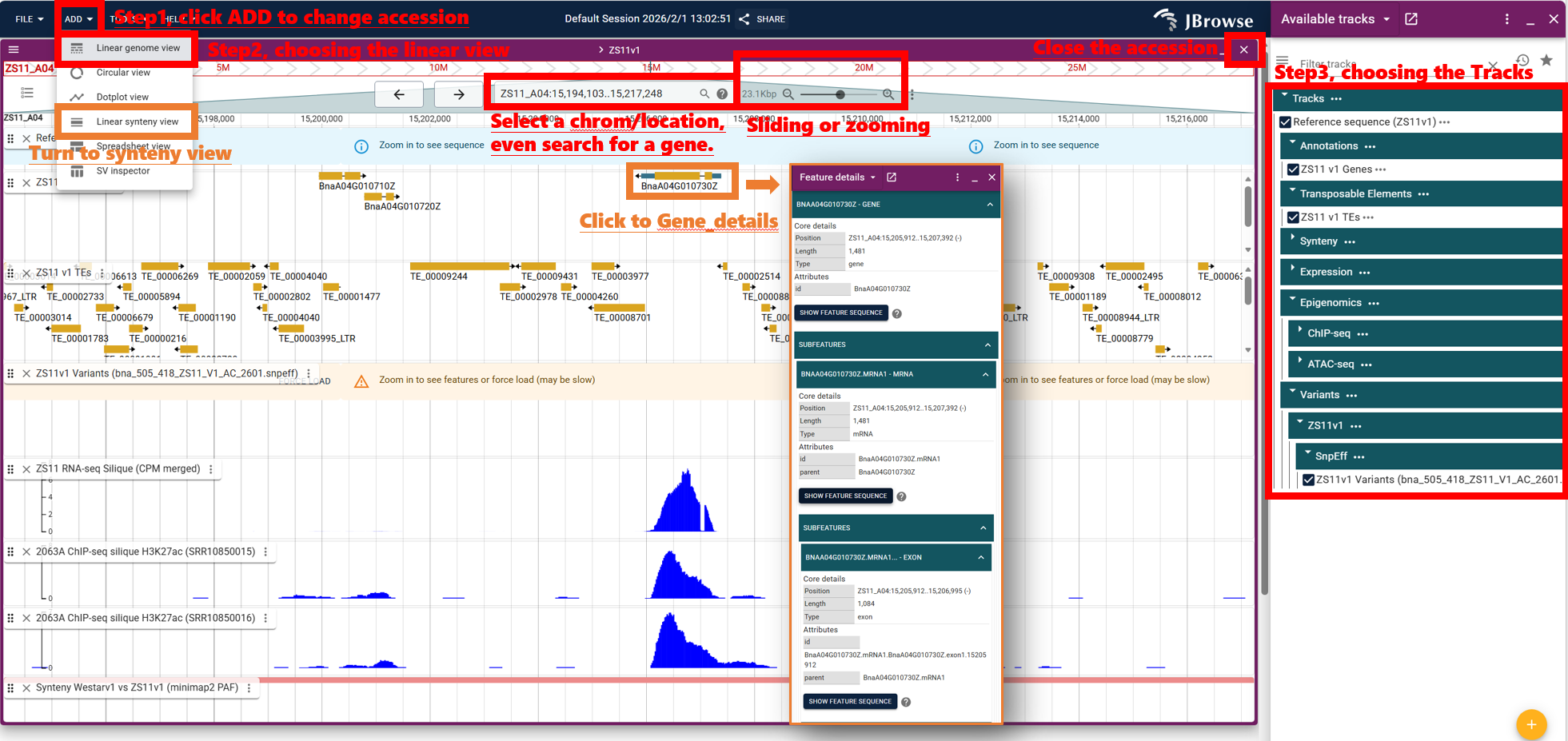
1) Choose an Assembly
- Click ADD (Linear genome view)to change accession
2) Turn on Tracks
- Open the Track Selector and enable the Gene track (and other tracks of interest, e.g. Synteny, Expression).
3) Search & Navigate
- Search by gene ID or jump to a region like chr:start-end.
- Click a gene to open details; expand subfeatures to inspect isoforms and sequences.
4) Compare Using Synteny
- Use the Linear synteny view or Dotplot view to follow collinear blocks and jump between corresponding loci.
For more information, click here to check official tutorial:
JBrowse2_Docs
Diesh et al, 2023. JBrowse 2: a modular genome browser with views of synteny and structural variation. Genome Biology 24:74.
https://doi.org/10.1186/s13059-023-02914-z